Buena Muerte Social Club#
Note
This is the companion notebook to the article Buena Muerte Social Club, submitted to FUN 2026. It is NOT intended to be self-contained. Please refer to the paper for context and explanations.
Preliminaries#
Basic Blood Types#
First, we define the different blood types. In addition to the \(2^3\) main types depending on the presence of \(A\), \(B\), and \(Rhesus\) markers, we add a void type that will be use later to model non-ghoul humans.
An important property is can_feed, that tells if a vampire of given type can feed on a ghoul of other type.
[1]:
import numpy as np
class BloodType:
A = ["", "A"]
B = ["", "B"]
R = ["-", "+"]
def __init__(self, i, j, k):
self.markers = np.array([i, j, k])
self.x = False
def __repr__(self):
if self.x:
return 'X'
i, j, k = self.markers
result = f"{self.A[i]}{self.B[j]}{self.R[k]}"
if len(result) == 1:
return f"O{result}"
else:
return result
@property
def sanitized_name(self):
res = self.__repr__()
return res.replace('-','m').replace('+','p')
def can_feed_on(self, other):
return np.all(other.markers <= self.markers)
@classmethod
def void(cls):
res = cls(1, 1, 1)
res.x = True
return res
Let us compute all possible groups. We’ll store them in a list, a class, and a dict for easy access.
[2]:
class Groups:
pass
groups = [BloodType(i, j, k) for i in range(2) for j in range(2) for k in range(2)]
g = Groups
for t in groups:
setattr(g, t.sanitized_name, t)
groups_ = {str(g): g for g in groups}
groups
[2]:
[O-, O+, B-, B+, A-, A+, AB-, AB+]
Vampire/Ghoul Types#
We now define Vampire/Ghoul types, which are defined by two blood types. The following properties are defined :
autarkic: the vampire can feed on their ghoul.
compatible_with: each vampire of two pairs can feed on the other one ghoul.
swinger: a non plug-and-drink pair compatible with another non plug-and-drink pair.
helpless: a non plug-and-drink pair that is only compatible with plug-and-drink-pairs.
three_compatible: each vampire of three pairs can feed on a distinct available ghoul, in a cyclic fashion.
[3]:
class VGType:
def __init__(self, v, g):
self.v = v
self.g = g
self.compatible_types = []
def __repr__(self):
return f"{self.v}/{self.g}"
@property
def category(self):
if self.auto_compatible:
return "autarkic"
if all(c.auto_compatible for c in self.compatible_types):
return "helpless"
return "swinger"
@property
def auto_compatible(self):
return self.v.can_feed_on(self.g)
def compatible_with(self, other):
return self.v.can_feed_on(other.g) and other.v.can_feed_on(self.g)
def three_cycle(self, other1, other2):
return self.v.can_feed_on(other1.g) and other1.v.can_feed_on(other2.g) and other2.v.can_feed_on(self.g)
def three_compatible_with(self, other1, other2):
return self.three_cycle(other1, other2) or self.three_cycle(other2, other1)
We now prepare all possible VG pairs and observe their repartition.
[4]:
from collections import Counter
vgs = [VGType(v, g) for v in groups for g in groups]
for v1 in vgs:
for v2 in vgs:
if v1.compatible_with(v2):
v1.compatible_types.append(v2)
vgs_ = {str(vg): i for i, vg in enumerate(vgs)}
Counter(vg.category for vg in vgs)
[4]:
Counter({'autarkic': 27, 'helpless': 19, 'swinger': 18})
Human types#
We call human a potential ghoul not affiliated yet to any vampire. Humans can be modeled as a VG pair where the vampire is a placeholder, denoted \(X\), compatible with everyone, e.g. X/O+
[5]:
class Human(VGType):
category = "human"
def __init__(self, g):
self.v = BloodType.void() # alway compatible
self.g = g
[6]:
humans = [Human(g) for g in groups]
humans
[6]:
[X/O-, X/O+, X/B-, X/B+, X/A-, X/A+, X/AB-, X/AB+]
Blood type distribution#
For numeric computation, we need statistics on the blood type probabilities. We leverage Wikipedia to access recent data.
[7]:
import requests
import re
from bs4 import BeautifulSoup as bs
from fake_useragent import UserAgent
ua = UserAgent()
soup = bs(requests.get("https://en.wikipedia.org/wiki/Blood_type_distribution_by_country", headers={'User-Agent': ua.random}).content)
table = soup('table')[1]('tr')
names = [s.text.strip().replace("−", "-") for s in table[0]('th')[2:]]
blood_type_by_country = dict()
for line in table[1:]:
country = line.th.text.strip().split('[')[0]
stats = {n: float(s.text.strip()[:-1]) if s.text[0].isdigit() else 0.0 for n, s in zip(names, line('td')[1:])}
total = sum(stats.values())
blood_type_by_country[country] = {k: v/total for k, v in stats.items()}
blood_type_by_country[country]['X'] = 1.0
[8]:
blood_type_by_country['United States']
[8]:
{'O+': 0.374,
'A+': 0.35700000000000004,
'B+': 0.085,
'AB+': 0.034,
'O-': 0.066,
'A-': 0.063,
'B-': 0.015,
'AB-': 0.006,
'X': 1.0}
[9]:
blood_type_by_country['France']
[9]:
{'O+': 0.365,
'A+': 0.382,
'B+': 0.077,
'AB+': 0.025,
'O-': 0.065,
'A-': 0.068,
'B-': 0.013999999999999999,
'AB-': 0.004,
'X': 1.0}
[10]:
blood_type_by_country['World']
[10]:
{'O+': 0.3878396121603878,
'A+': 0.2757297242702757,
'B+': 0.0818099181900818,
'AB+': 0.0201999798000202,
'O-': 0.1323098676901323,
'A-': 0.0818099181900818,
'B-': 0.0201999798000202,
'AB-': 0.00010099989900010099,
'X': 1.0}
Given a list of VG types and a country, issue normalized arrival rates.
[11]:
def arrival_rates(vgs, country="United States"):
probs = blood_type_by_country[country]
res = [probs[str(vg.v)]*probs[str(vg.g)] for vg in vgs ]
total = sum(res)
return [r/total for r in res]
For the record, one can compute distribution per VG category:
[12]:
from collections import defaultdict
shares = defaultdict(float)
rates = arrival_rates(vgs)
for i, vg in enumerate(vgs):
shares[vg.category] += float(100*rates[i])
dict(shares)
[12]:
{'autarkic': 56.6078, 'helpless': 28.1786, 'swinger': 15.2136}
Instability study#
We know that the base model is not stabilisable. Yet, we still can investigate through simulations the practical performance of robust matching algorithm in that settings.
Towards that goal, we write a function that converts a list of VG types into a matching problem.
[13]:
import stochastic_matching as sm
def build_model(vgs, country="United States", self=True, relaxed=False, threesomes=False):
edges = []
n = len(vgs)
for i, vg in enumerate(vgs):
if relaxed or (self and vg.auto_compatible):
edges.append([i])
for j in range(i+1, n):
if vg.compatible_with(vgs[j]):
edges.append([i, j])
if not threesomes:
continue
for k in range(j+1, n):
if vg.three_compatible_with(vgs[j], vgs[k]):
if any(str(p.v)==str(p.g) for p in [vg, vgs[j], vgs[k]]):
continue
edges.append([i, j, k])
incidence = np.zeros((n, len(edges)), dtype=int)
for i, e in enumerate(edges):
incidence[e, i] = 1
model = sm.Model(incidence=incidence, names=[str(vg) for vg in vgs],
rates=arrival_rates(vgs, country=country))
model.cats = [vg.category for vg in vgs]
model.edges = edges
try:
model.adjacency = sm.model.incidence_to_adjacency(model.incidence)
except ValueError:
pass
return model
Swingers-only#
We first focus on the inter-dependent types, because it is smaller and makes sense from a selfish perspective.
[14]:
swingers = [vg for vg in vgs if vg.category == 'swinger']
model = build_model(swingers)
model.show_graph()
Nice structure.
[15]:
model.kernel
[15]:
Kernels of a graph with 18 nodes and 24 edges.
Node dimension is 0.
Edge dimension is 6
Type: Surjective-only
Node kernel:
[]
Edge kernel:
[[ 0.17028994 -0.17028994 -0.07412584 0.07412584 0. -0.17028994
-0.12829445 0.12829445 0. 0.07412584 0.12829445 0.
-0.32576712 0.19747267 -0.44229877 0.36817293 0.19747267 -0.19747267
0.36817293 -0.36817293 0.12771746 0.04257249 0.04257249 -0.04257249]
[ 0.04439801 -0.04439801 -0.26529933 0.26529933 0. -0.04439801
-0.33257364 0.33257364 0. 0.26529933 0.33257364 0.
-0.38079546 0.04822182 0.0484483 -0.31374763 0.04822182 -0.04822182
-0.31374763 0.31374763 0.03329851 0.0110995 0.0110995 -0.0110995 ]
[ 0.35072029 -0.35072029 0.12045588 -0.12045588 0. -0.35072029
-0.113703 0.113703 0. -0.12045588 0.113703 0.
0.12788313 -0.24158613 0.12232013 -0.00186425 -0.24158613 0.24158613
-0.00186425 0.00186425 0.51304022 -0.16231993 -0.16231993 0.16231993]
[ 0.02262008 -0.02262008 -0.20995298 0.20995298 0. -0.02262008
-0.08260737 0.08260737 0. 0.20995298 0.08260737 0.
0.28638173 -0.3689891 -0.29147499 0.08152201 -0.3689891 0.3689891
0.08152201 -0.08152201 -0.23303494 0.25565502 0.25565502 -0.25565502]
[ 0.33486514 -0.33486514 -0.03178975 0.03178975 0. -0.33486514
0.22875339 -0.22875339 0. 0.03178975 -0.22875339 0.
0.05848433 0.17026906 0.11609151 -0.14788126 0.17026906 -0.17026906
-0.14788126 0.14788126 0.00114885 0.33371628 0.33371628 -0.33371628]
[ 0.00676492 -0.00676492 -0.36219861 0.36219861 0. -0.00676492
0.25984901 -0.25984901 0. 0.36219861 -0.25984901 0.
0.21698293 0.04286608 -0.29770361 -0.064495 0.04286608 -0.04286608
-0.064495 0.064495 0.25507369 -0.24830877 -0.24830877 0.24830877]]
As predicted, it is not stabilizable.
[16]:
model.stabilizable
[16]:
False
[17]:
def grouped_delay(simu):
from collections import defaultdict
import numpy as np
wait = defaultdict(float)
for i, cat in enumerate(simu.model.cats):
wait[cat] += float(simu.avg_queues[i])
return dict(wait)
[18]:
base_params = {"simulator": "longest", "n_steps": 10**9, "max_queue": 100000, "seed": 42}
[19]:
xp = sm.XP("Swingers", model=model, **base_params)
res = sm.evaluate(xp, [grouped_delay])
res
[19]:
{'Swingers': {'grouped_delay': {'swinger': 43048.157124944}}}
Helpless-excluded#
Exluding helpless types is harsh but leads to strong stability.
We first remove the self-loops to have a simple graph to display (it is only for display).
[45]:
helpfuls = [vg for vg in vgs if vg.category != 'helpless']
model = build_model(helpfuls, self=False)
colors = {"autarkic": "#00FF00", "swinger": "#FF8888", "helpless": "#FF0000"}
physics = {
"solver": "barnesHut",
"barnesHut": {
"gravitationalConstant": -15000,
"centralGravity": 0.25,
"springLength": 200,
"springConstant": 0.0015,
"damping": 0.095,
"avoidOverlap": 0.12
},
"maxVelocity": 15,
"stabilization": {
"iterations": 800,
"updateInterval": 25
}
}
nodes_info = [{"color": {"background": colors[vg.category]}} for vg in helpfuls]
options = {"width": "1000px", "nodes": {"font": {"size": 60}}, "physics": physics}
model.show_graph(vis_options=options, nodes_info=nodes_info)
Back to the real thing.
[21]:
model = build_model(helpfuls)
model.kernel
[21]:
Kernels of a graph with 45 nodes and 317 edges.
Node dimension is 0.
Edge dimension is 272
Type: Surjective-only
Node kernel:
[]
Edge kernel:
[[ 1.85577352e-01 4.60120460e-02 7.29821545e-02 ... 4.78644736e-04
-1.19916425e-03 -1.67780899e-03]
[ 1.70012321e-01 2.91401361e-02 6.69110172e-02 ... 2.85079976e-03
9.21133624e-04 -1.92966614e-03]
[ 1.51967077e-01 2.44349889e-02 7.03749366e-02 ... -1.77408140e-03
-2.70700952e-04 1.50338044e-03]
...
[-2.89699459e-02 -1.53198499e-02 -1.07370205e-02 ... 9.29353881e-01
-6.42094593e-02 6.43665994e-03]
[ 8.74057828e-03 8.55991905e-04 3.83245243e-04 ... -6.30747725e-02
8.15041946e-01 -1.21883281e-01]
[ 3.77105242e-02 1.61758418e-02 1.11202657e-02 ... 7.57134673e-03
-1.20748594e-01 8.71680059e-01]]
[22]:
model.stabilizable
[22]:
True
[23]:
base_params["simulator"] = "virtual_queue"
xp = sm.XP("Helpfuls", model=model, **base_params)
res = sm.evaluate(xp, [grouped_delay])
res
[23]:
{'Helpfuls': {'grouped_delay': {'autarkic': 4.162427954,
'swinger': 10.752904849000002}}}
All categories#
We first remove the self-loops to have a simple graph to display (it is only for display).
[46]:
model = build_model(vgs, self=False)
nodes_info = [{"color": {"background": colors[vg.category]}} for vg in vgs]
options = {"width": "1000px", "nodes": {"font": {"size": 60}}, "physics": physics}
model.show_graph(vis_options=options, nodes_info=nodes_info)
Back to the real thing.
[25]:
model = build_model(vgs)
model.kernel
[25]:
Kernels of a graph with 64 nodes and 378 edges.
Node dimension is 0.
Edge dimension is 314
Type: Surjective-only
Node kernel:
[]
Edge kernel:
[[-8.07660786e-02 -5.14815963e-03 1.55185186e-01 ... 1.23681204e-03
1.07651019e-03 -1.60301853e-04]
[-5.27865346e-02 1.35874032e-02 1.66497191e-01 ... 4.31476721e-03
3.36397446e-03 -9.50792748e-04]
[-7.31481008e-02 -1.48678109e-02 1.68269311e-01 ... -8.36046328e-04
1.31250595e-03 2.14855228e-03]
...
[ 1.05249784e-02 1.28211539e-02 -4.16977264e-02 ... 9.33214398e-01
-5.95732615e-02 7.21234047e-03]
[ 3.51189245e-02 1.37009886e-02 -1.00579058e-01 ... -5.94140329e-02
8.18611345e-01 -1.21974622e-01]
[ 2.45939461e-02 8.79834759e-04 -5.88813319e-02 ... 7.37156906e-03
-1.21815393e-01 8.70813038e-01]]
[26]:
model.stabilizable
[26]:
False
[27]:
base_params["simulator"] = "virtual_queue"
xp = sm.XP("All categories", model=model, **base_params)
res = sm.evaluate(xp, [grouped_delay])
res
[27]:
{'All categories': {'grouped_delay': {'autarkic': 0.0011420769999999998,
'helpless': 49526.04985507099,
'swinger': 10186.265578822002}}}
Threesomes#
[28]:
three = build_model(vgs, threesomes=True)
three.kernel
[28]:
Kernels of a graph with 64 nodes and 4397 edges.
Node dimension is 0.
Edge dimension is 4333
Type: Surjective-only
Node kernel:
[]
Edge kernel:
[[ 3.89524050e-02 -9.99891026e-02 1.02077254e-02 ... 3.03768858e-04
1.54222108e-04 -1.49546751e-04]
[ 4.03133967e-02 -1.01467934e-01 1.12084971e-02 ... -3.78626371e-05
-1.29415410e-04 -9.15527730e-05]
[ 4.00204502e-02 -1.01827540e-01 1.07834303e-02 ... -3.54594281e-05
-1.62044872e-04 -1.26585444e-04]
...
[ 2.34574620e-03 -6.13999101e-03 5.82162293e-04 ... 9.96407855e-01
-3.22429376e-03 3.67850837e-04]
[-1.13259079e-02 2.88977694e-02 -3.03025921e-03 ... -3.23536900e-03
8.71888904e-01 -1.24875727e-01]
[-1.36716541e-02 3.50377604e-02 -3.61242151e-03 ... 3.56775597e-04
-1.24886802e-01 8.74756422e-01]]
[29]:
xp = sm.XP("threesomes allowed", model=three, **base_params)
res = sm.evaluate(xp, [grouped_delay])
res
[29]:
{'threesomes allowed': {'grouped_delay': {'autarkic': 265.46645575099996,
'helpless': 21061.442134908,
'swinger': 7806.321084564999}}}
Stairway to Stability#
Safety valves#
[30]:
model = build_model(vgs, relaxed=True)
[31]:
model.kernel
[31]:
Kernels of a graph with 64 nodes and 415 edges.
Node dimension is 0.
Edge dimension is 351
Type: Surjective-only
Node kernel:
[]
Edge kernel:
[[-6.66392973e-02 1.46326644e-01 -7.22319972e-03 ... -3.38870783e-04
-1.48396032e-03 -1.14508953e-03]
[-9.99357307e-02 1.29482979e-01 -1.12844994e-02 ... -1.31766036e-03
-2.37050539e-03 -1.05284503e-03]
[-2.67131450e-02 1.58095770e-01 -2.74271795e-03 ... -3.87754635e-04
-1.69770748e-03 -1.30995284e-03]
...
[ 1.23332381e-02 -3.49292348e-02 -3.68281453e-03 ... 9.33285059e-01
-6.02510432e-02 6.46389731e-03]
[-4.70729024e-03 1.05815809e-02 -7.12851020e-04 ... -5.92526383e-02
8.18992064e-01 -1.21755297e-01]
[-1.70405283e-02 4.55108156e-02 2.96996352e-03 ... 7.46230223e-03
-1.20756893e-01 8.71780805e-01]]
[32]:
model.stabilizable
[32]:
True
[33]:
base_params['max_queue'] = 20000
forbidden_edges = [i for i, e in enumerate(model.edges)
if len(e)==1 and not vgs[e[0]].auto_compatible]
xp = sm.XP('discarding', model=model,
iterator=sm.Iterator('k', [2**i for i in range(1, 12)]),
forbidden_edges=forbidden_edges, **base_params)
[34]:
import multiprocess as mp
with mp.Pool() as p:
res = sm.evaluate(xp, ["regret", grouped_delay], pool=p)
[35]:
from matplotlib import pyplot as plt
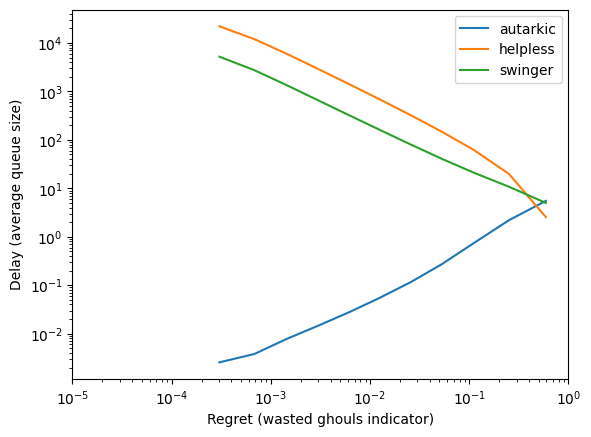
r = res['discarding']
x = r['regret']
ys = r['grouped_delay']
for l in ys[0].keys():
y = [yy[l] for yy in ys]
plt.loglog(x, y, label=l)
plt.xlabel('Regret (wasted ghouls indicator)')
plt.ylabel('Delay (average queue size)')
plt.xlim([.00001, 1])
plt.legend()
import tikzplotlib
tikzplotlib.save("waste.tex")
plt.show()
Humans#
We now incoporate some amount of humans (vampire-free pre-ghouls) into the mix.
[36]:
def mix_model(α = .001):
μc = arrival_rates(vgs)
μd = arrival_rates(humans)
μ = [(1-α)*p/np.sum(μc) for p in μc]+[α*p/np.sum(μd) for p in μd]
model = build_model(vgs+humans)
model.rates = μ
return model
Note
If we try to numerically compute the minimum proportion of humans that allows the system to be stable, one gets a positive value. This is due to the numerical margin of error of the best positive position. For highly skewed problem, it can be difficult to distinguish a zeroed edge from a barely positive one.
Yet, the value found has its one interest: it gives us a bound below which stability may be difficult to observe.
[37]:
def find_threshold(steps=20):
u = 1.0
l = 0.0
for _ in range(steps):
c = (u+l)/2
m = mix_model(c)
if m.stabilizable:
u = c
else:
l = c
return (u+l)/2
find_threshold()
[37]:
0.0004916191101074219
[38]:
base_params['simulator'] = 'virtual_queue'
xp = sm.XP('humanities', iterator=sm.Iterator('model', [2**i*.0005 for i in range(11)], 'alpha', lambda a: mix_model(a)), **base_params)
[39]:
import multiprocess as mp
with mp.Pool() as p:
res = sm.evaluate(xp, [grouped_delay], pool=p)
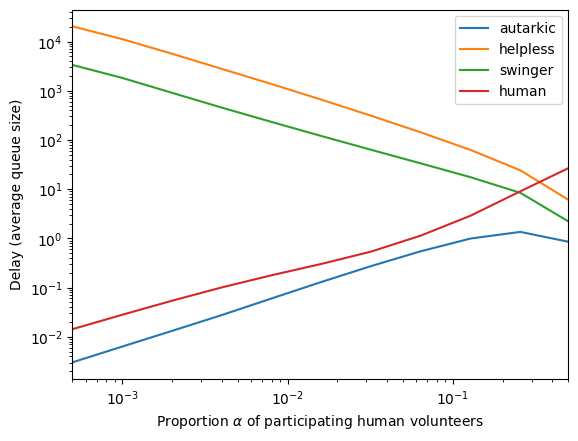
[40]:
r = res['humanities']
x = r['alpha']
ys = r['grouped_delay']
for l in ys[0].keys():
y = [e[l] for e in ys]
plt.loglog(x, y, label=l)
plt.xlabel('Proportion $\\alpha$ of participating human volunteers')
plt.ylabel('Delay (average queue size)')
plt.xlim([.0005, .5])
plt.legend()
import tikzplotlib
tikzplotlib.save("humans.tex")
plt.show()